Tutorial 5: Parameter sweeps
A parameter sweep runs the same scenario across a range of input values so you can study how outcomes change with each parameter. This tutorial sweeps over treatment-seeking coverage to show how different levels of case management affect malaria burden in a seasonal setting.
File: tutorials/tutorial_5_sweep.py
SimulationBuilder
SimulationBuilder creates multiple simulations by combining sweep definitions. Each sweep
definition is a callback function paired with a list of values — the builder calls the
function once per value per simulation.
Adding two sweep definitions produces a cross-product: every combination of treatment_coverage and
Run_Number gets its own simulation. With 3 coverage values and 3 random seeds that is 9
simulations total:
builder = SimulationBuilder()
builder.add_sweep_definition(update_campaign, [0.3, 0.6, 0.9])
builder.add_sweep_definition(sweep_run_number, [0, 1, 2])
Sweeping campaign parameters
Coverage is now a parameter rather than a constant. update_campaign() is the sweep callback
that rebuilds the campaign for each simulation with the assigned coverage value:
partial() binds treatment_coverage to build_campaign so the resulting function takes no arguments,
which is what create_campaign_from_callback() expects. The dict returned becomes a tag on
the simulation — used below to group results by coverage value.
build_campaign() accepts treatment_coverage as a parameter and applies it to both clinical and severe
case targets:
ITN coverage remains fixed at 0.5 — only treatment-seeking coverage is swept.
Grouping results by coverage
After downloading, group_by_coverage() reads the treatment_coverage tag from each simulation in
the experiment object and moves its downloaded directory into a subdirectory named by coverage
value:
Reading tags from the experiment object works on all platforms — Container, COMPS, and SLURM.
After grouping, tutorial_5_results/ contains one subdirectory per coverage value:
tutorial_5_results/
treatment_coverage_0.3/
{sim_id}/InsetChart.json ← Run_Number 0
{sim_id}/InsetChart.json ← Run_Number 1
{sim_id}/InsetChart.json ← Run_Number 2
treatment_coverage_0.6/
...
treatment_coverage_0.9/
...
Plotting by coverage group
plot_mean() takes one directory per coverage group and uses the directory name as the legend
label. show_raw_data=True overlays the individual simulation lines in a lighter color so
the stochastic spread within each group is visible alongside the mean:
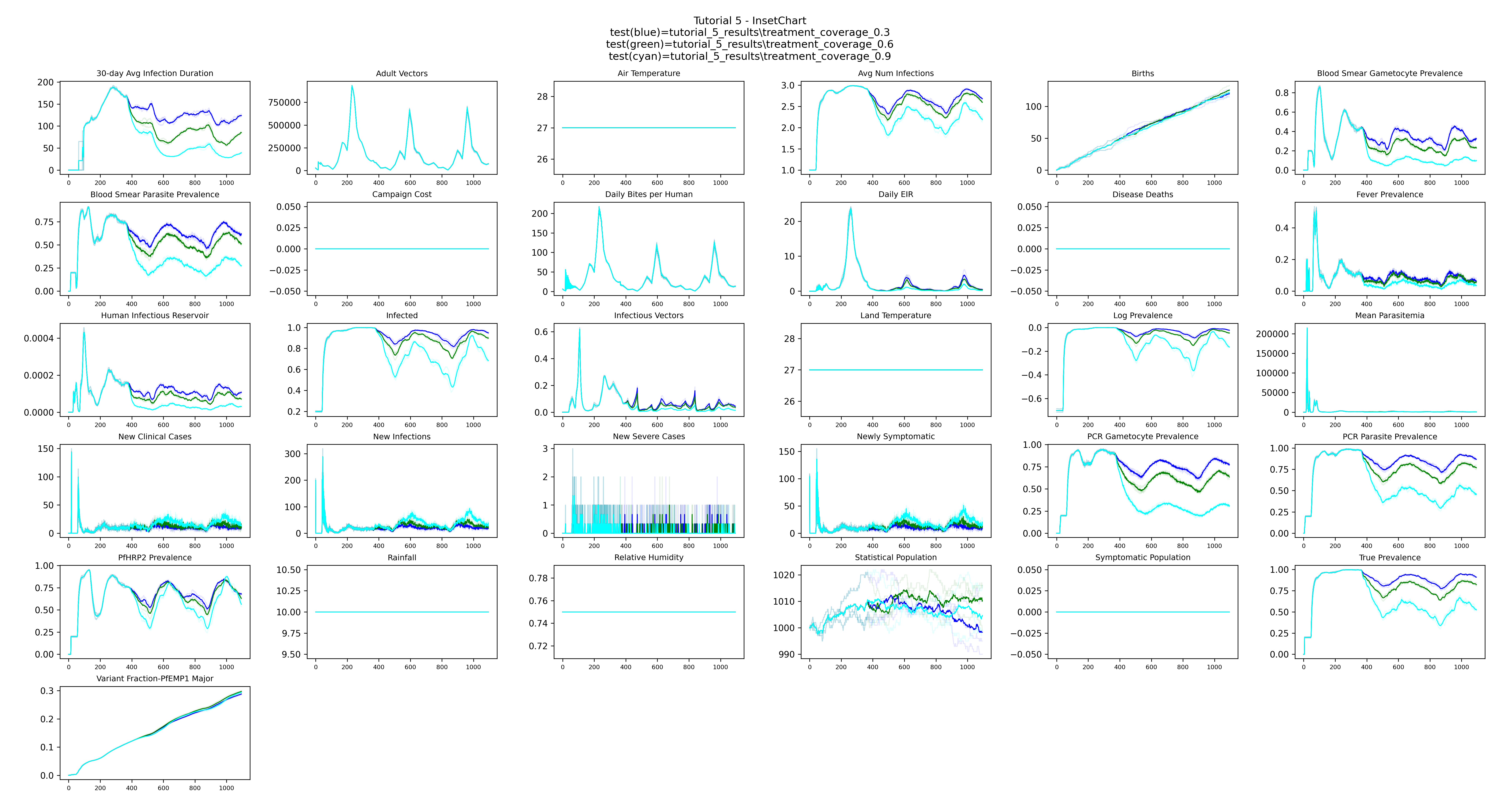
Example output
The plot shows one bold mean line per coverage group (treatment_coverage_0.3, treatment_coverage_0.6, treatment_coverage_0.9), with the
three individual stochastic runs shown in a lighter color behind each mean. With higher
coverage, there are fewer infected people (Infected channel) and each infection resolves more
quickly (30-day Avg Infection Duration). Together, fewer infected people spending less time
infected means mosquitoes have less opportunity to acquire infection from humans — resulting
in fewer infectious vectors and a lower Daily EIR.
Next
Tutorial 6 introduces calibration with CalibManager, fitting
x_Temporary_Larval_Habitat to match a reference PfPR target.