Gene Drives
Existing malaria control tools — insecticide-treated nets, indoor residual spraying, and antimalarial drugs — have driven significant reductions in transmission, but the spread of insecticide and drug resistance threatens to erode these gains. Reaching elimination in high-transmission settings will likely require new tools that can self-propagate through wild mosquito populations without continuous mass releases.
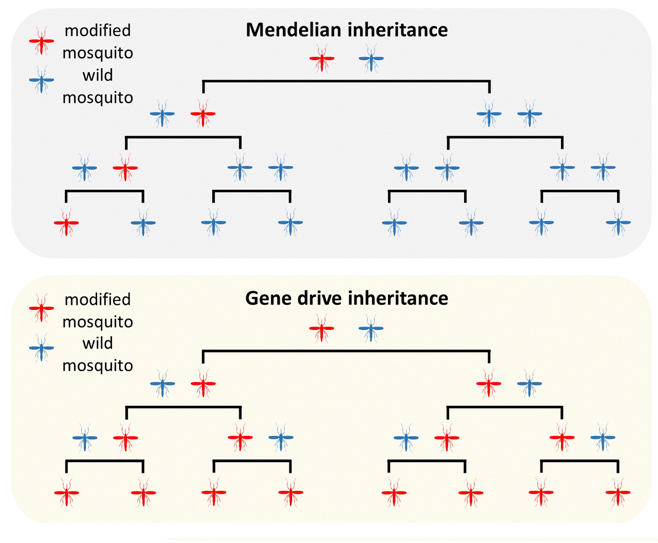
Gene drives are genetic elements that bias their own inheritance beyond what Mendelian genetics predicts. In a standard heterozygote, a transgene has a 50% chance of being passed to each offspring and is likely lost over time if it carries any fitness cost. A gene drive overcomes this: most or all offspring of a heterozygous carrier inherit the drive element, allowing it to spread rapidly to high frequency within a population over just a few generations (Figure 1).
Figure 1. Mendelian inheritance (top) versus gene drive inheritance (bottom). Under Mendelian inheritance, a modified mosquito (red) mated with a wild mosquito (blue) produces 50% modified offspring, so the modification stays rare or is lost. A gene drive causes most or all offspring to inherit the drive element, enabling rapid population-wide spread. From Hammond and Galizi (2017), doi:10.1080/20477724.2018.1438880.
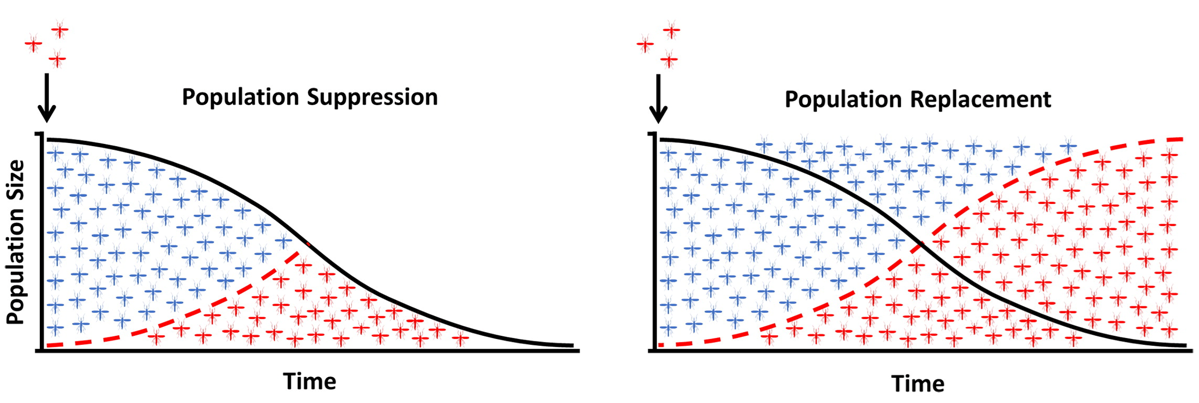
Two broad strategies have been proposed for gene-drive-based malaria control (Figure 2). Population suppression drives are designed to reduce mosquito reproductive capacity — for example, by distorting the sex ratio toward males — eventually reducing the population below the threshold required to sustain transmission. Population replacement drives spread a refractory trait (inability to transmit malaria) through the population, leaving mosquito numbers intact but eliminating their capacity to transmit disease.
Figure 2. Population suppression (left) reduces the total vector population over time. Population replacement (right) maintains population size but converts wild mosquitoes (blue) to modified, malaria-refractory mosquitoes (red). From Hammond and Galizi (2017), doi:10.1080/20477724.2018.1438880.
A further distinction is between self-sustaining drives, which are designed to spread indefinitely once released, and self-limiting drives, which are engineered to dissipate over time — an important consideration for ecological containment and regulatory approval.
EMOD supports five gene drive types that model these different strategies. Gene drives
are configured in the Drivers array within each species entry in Vector_Species_Params.
Note
The model does not allow for mixing drive types within a species.
Seealso
Hammond, A.M. and Galizi, R. (2017). Gene drives to fight malaria: current state and future directions. Pathogens and Global Health, 111(8), 412–423.
Leung, S., Windbichler, N., Wenger, E.A., Bever, C.A. and Selvaraj, P. (2022). Population replacement gene drive characteristics for malaria elimination in a range of seasonal transmission settings: a modelling study. Malaria Journal, 21, 226.
Vitale, M., Kranjc, N., Leigh, J., Kyrou, K., Courty, T., Marston, L., Grilli, S., Crisanti, A. and Bernardini, F. (2024). Y chromosome shredding in Anopheles gambiae: insight into the cellular dynamics of a novel synthetic sex ratio distorter. PLOS Genetics, 20(6), e1011303.
CLASSIC
The classic gene drive bundles a Cas9 endonuclease and a guide RNA (gRNA) at the same locus as
the drive allele (represented by Driving_Allele). When two gametes come together during
genome formation, if one gamete has the driving allele and the other has the target allele
(Allele_To_Replace), the drive cuts the target and copies the driving allele into the cut
site.
The drive requires all of the following conditions to be met:
- The
Driving_Allelemust be present in exactly one of the two gametes (not both). - The
Allele_To_Replacefor the driving allele must exist in the other gamete. - The copy of the driving allele itself must succeed (based on
Copy_To_Likelihood).
If any of these conditions is not met, the entire drive fails and standard Mendelian inheritance
applies. When driving additional loci (effectors), the Allele_To_Copy must exist in the
gamete with the driving allele, and the Allele_To_Replace must exist in the other gamete.
When the conversion is not perfect, you can configure the drive to model different outcomes at the cut site:
- Successful copy: The drive allele is copied (super-Mendelian inheritance).
- Resistance allele: Non-homologous end joining (NHEJ) creates a resistant allele that can no longer be cut, preventing future drive conversion at that locus.
- No change: The cut fails entirely.
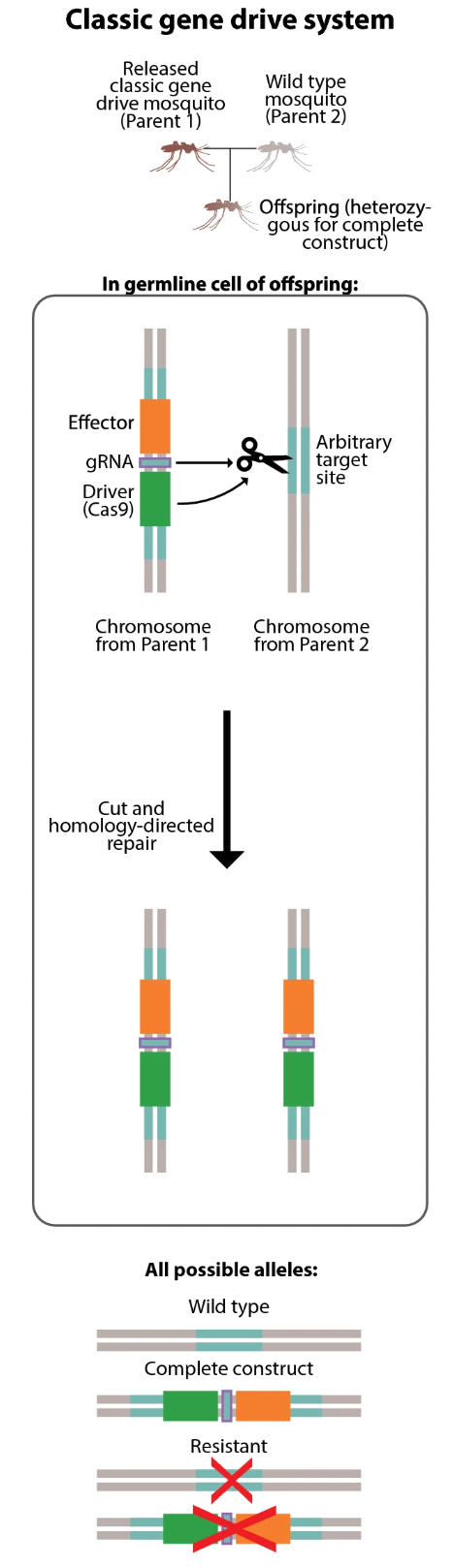
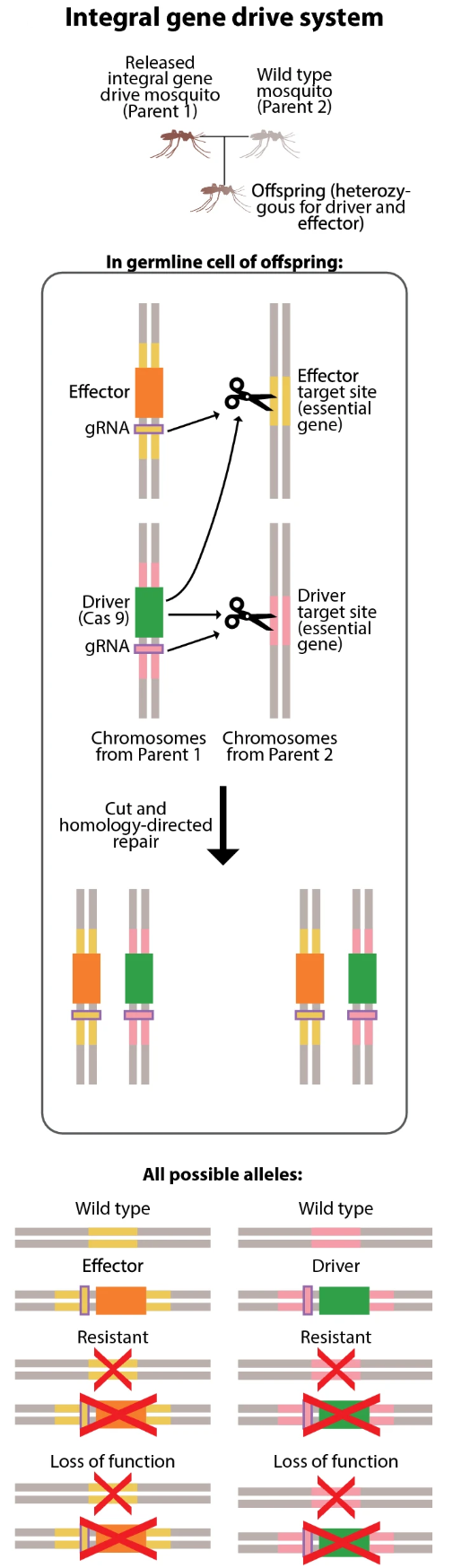
Classic gene drive system. A drive mosquito mates with a wild-type mosquito; in the offspring germline, the Cas9 and gRNA cut the wild-type chromosome at the target site and homology-directed repair copies the complete construct. Possible alleles in offspring: wild type (no copy), complete construct (successful copy), or resistant (non-homologous end joining creates a mutated target site the drive can no longer recognize). From Leung et al. (2022), doi:10.1186/s12936-022-04242-2.
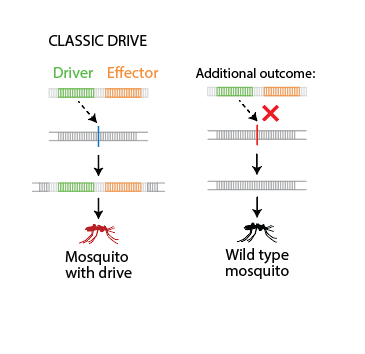
In EMOD, this mechanism is abstracted as two alleles — a Driver (representing Cas9) and an
Effector — bundled at the same locus. The drive either copies successfully or fails, as shown
below; these outcomes map directly to the probabilities set in Copy_To_Likelihood.
EMOD abstraction of a classic gene drive: the Driver and Effector are bundled at the same locus. On the left, the drive copies successfully into the target chromosome, producing a mosquito carrying the drive. On the right, the additional outcome where the drive fails and the wild-type allele is retained.
Example configuration
A classic drive at a single locus. The Ade allele bundles Cas9 + gRNA and replaces the
wild-type allele Aw with 99% efficiency. There is a 0.7% chance of complete failure where
the wild-type allele is retained, and a 0.3% chance of a mutation to Am — a resistant allele
that the drive can no longer recognize, preventing future conversion at that locus:
{
"Drivers": [
{
"Driver_Type": "CLASSIC",
"Driving_Allele": "Ade",
"Alleles_Driven": [
{
"Allele_To_Copy": "Ade",
"Allele_To_Replace": "Aw",
"Copy_To_Likelihood": [
{"Copy_To_Allele": "Aw", "Likelihood": 0.007},
{"Copy_To_Allele": "Ade", "Likelihood": 0.990},
{"Copy_To_Allele": "Am", "Likelihood": 0.003}
]
}
]
}
]
}
INTEGRAL_AUTONOMOUS
The integral autonomous drive separates the Cas9 (driver) and gRNA (effector) onto different loci. Unlike the classic drive, it can drive the effector even when the driver allele itself fails to copy. It can copy an effector allele from either gamete into the other, since the Cas9 produced by the driver acts across gametes regardless of which gamete carries the driver.
The drive activates whenever at least one copy of the driver allele is present in the genome —
it does not require heterozygosity at the driven locus. For each driven locus, the
Allele_To_Copy must exist in the gamete with the driving allele, and the
Allele_To_Replace must exist in the other gamete. If one of these conditions is not met for
a particular locus, nothing happens at that locus, but other loci can still be driven.
Integral gene drive system. Driver and effector are on separate loci, each with their own gRNA, allowing each to be copied independently. Possible alleles at each locus: wild type, the introduced construct, resistant (drive can no longer recognize the target site), or loss-of-function (lethal mutation at an essential gene target site). From Leung et al. (2022), doi:10.1186/s12936-022-04242-2.
In EMOD, the Driver and Effector are modeled as alleles at separate loci. Because they are independent, each can succeed or fail on its own — producing a mosquito carrying both (most effective), only the effector, only the driver, or neither, as shown below.
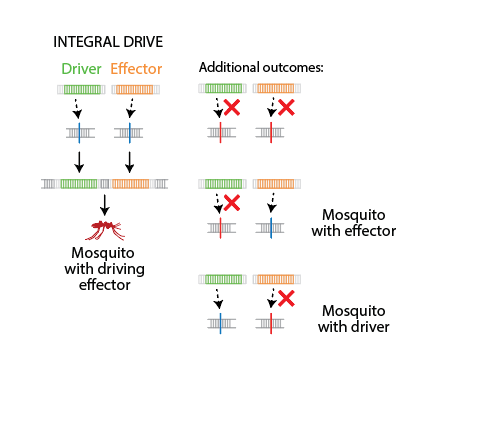
EMOD abstraction of an integral drive: the Driver and Effector are at separate loci and copy independently. The main outcome (left) is a mosquito carrying both. Additional outcomes (right) show the effector copying without the driver, the driver copying without the effector, or both failing.
Example configuration
Driver locus Ad replaces wild-type Aw with 90% efficiency (6% failure, 4% mutation to
resistant Am). Effector locus Be replaces wild-type Bw with 80% efficiency (15% failure,
5% mutation to resistant Bm). Because the loci are independent, the effector can be driven
even if the driver fails to copy, and vice versa:
{
"Drivers": [
{
"Driver_Type": "INTEGRAL_AUTONOMOUS",
"Driving_Allele": "Ad",
"Alleles_Driven": [
{
"Allele_To_Copy": "Ad",
"Allele_To_Replace": "Aw",
"Copy_To_Likelihood": [
{"Copy_To_Allele": "Aw", "Likelihood": 0.06},
{"Copy_To_Allele": "Ad", "Likelihood": 0.90},
{"Copy_To_Allele": "Am", "Likelihood": 0.04}
]
},
{
"Allele_To_Copy": "Be",
"Allele_To_Replace": "Bw",
"Copy_To_Likelihood": [
{"Copy_To_Allele": "Bw", "Likelihood": 0.15},
{"Copy_To_Allele": "Be", "Likelihood": 0.80},
{"Copy_To_Allele": "Bm", "Likelihood": 0.05}
]
}
]
}
]
}
DAISY_CHAIN
A daisy chain drive is a multi-element system where each component drives the next in a chain, but the first element in the chain has no drive acting on it. This creates a self-limiting drive: the first element is lost from the population through standard Mendelian dilution, which eventually removes the driving force from downstream elements. Daisy chain drives are modeled as a variant of the autonomous drive with specific locus dependencies.
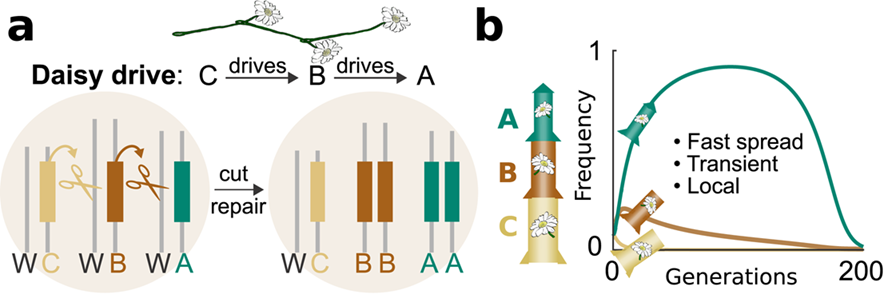
Daisy chain drive. (a) CRISPR components are separated so each element drives the next: C drives B, B drives A. C is not driven and is lost in half of offspring; once C is gone, B loses its driver and is lost in turn, continuing until the drive stops. (b) The loss of non-driving elements is analogous to gravity on a rocket — adding more elements allows the system to spread further before it runs out of genetic fuel. From Esvelt and Gemmell (2017), doi:10.1371/journal.pbio.2003850.
Example configuration
A two-element daisy chain. Ct (at the end of the chain) drives Bt into the population.
Bt (in the middle) drives At, the effector. Nothing drives Ct itself, so it is lost
through Mendelian dilution over time, making the drive self-limiting.
Note
Because DAISY_CHAIN is implemented as a variant of INTEGRAL_AUTONOMOUS, Alleles_Driven
must include an entry for the driver allele's own locus. This entry defines what happens
at that locus when the drive is active. For a daisy element that does not drive itself,
set the Copy_To_Likelihood for that entry to 100% failure (Likelihood: 1.0 for
the Allele_To_Replace), as shown below for Bt and Ct.
{
"Drivers": [
{
"Driver_Type": "DAISY_CHAIN",
"Driving_Allele": "Bt",
"Alleles_Driven": [
{
"Allele_To_Copy": "At",
"Allele_To_Replace": "Aw",
"Copy_To_Likelihood": [
{"Copy_To_Allele": "Aw", "Likelihood": 0.0},
{"Copy_To_Allele": "At", "Likelihood": 1.0}
]
},
{
"Allele_To_Copy": "Bt",
"Allele_To_Replace": "Bw",
"Copy_To_Likelihood": [
{"Copy_To_Allele": "Bw", "Likelihood": 1.0}
]
}
]
},
{
"Driver_Type": "DAISY_CHAIN",
"Driving_Allele": "Ct",
"Alleles_Driven": [
{
"Allele_To_Copy": "Bt",
"Allele_To_Replace": "Bw",
"Copy_To_Likelihood": [
{"Copy_To_Allele": "Bw", "Likelihood": 0.0},
{"Copy_To_Allele": "Bt", "Likelihood": 1.0}
]
},
{
"Allele_To_Copy": "Ct",
"Allele_To_Replace": "Cw",
"Copy_To_Likelihood": [
{"Copy_To_Allele": "Cw", "Likelihood": 1.0}
]
}
]
}
]
}
X_SHRED and Y_SHRED
Sex-ratio distortion drives work by destroying (shredding) sex-chromosome-bearing gametes during
spermatogenesis. These drives only apply to gametes created from a male genome and are configured
via the Shredding_Alleles parameter block rather than Alleles_Driven.
- X_SHRED: Destroys X-bearing sperm, biasing offspring toward males. Requires the male
parent to carry the driving allele (
Allele_Required) on the Y chromosome. - Y_SHRED: Destroys Y-bearing sperm, biasing offspring toward females. Requires the male
parent to carry the driving allele (
Allele_Required) on the X chromosome.
Each shredding drive specifies:
Allele_Required— the allele the male must carry for shredding to occurAllele_To_Shred— the sex-chromosome allele that is targeted for destructionAllele_To_Shred_To— the allele that surviving shredded gametes are converted toAllele_Shredding_Fraction— the fraction of targeted gametes destroyed (0.0--1.0, default 1.0)Allele_To_Shred_To_Surviving_Fraction— the fraction of shredded gametes that survive as theAllele_To_Shred_Toallele rather than being eliminated (0.0--1.0, default 0.0)
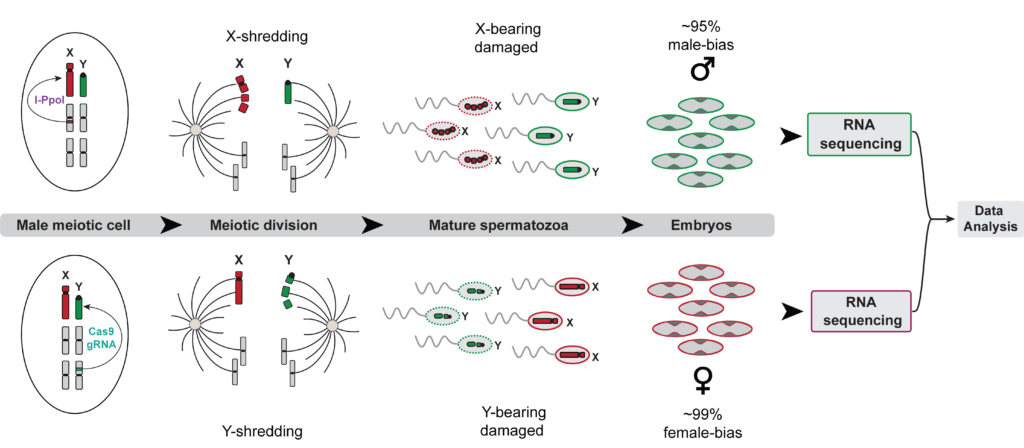
X-shredding (top) and Y-shredding (bottom) during male meiosis. In X-shredding, an endonuclease (I-PpoI) damages X-bearing sperm during meiotic division, leaving primarily Y-bearing sperm viable and producing approximately 95% male-biased offspring. In Y-shredding, a Cas9/gRNA construct targets and damages Y-bearing sperm, leaving primarily X-bearing sperm viable and producing approximately 99% female-biased offspring. From Vitale, M. (2024), Target Malaria, Y-chromosome shredding in Anopheles gambiae.
Example configuration
An X-shredding drive that biases offspring toward males. The drive allele Ad is at a
non-gender locus. When a male carrying Ad also has the Y-chromosome allele Yw
(Allele_Required), his X-bearing sperm carrying Xw (Allele_To_Shred) are destroyed.
Driving_Allele_Params specifies how the drive allele itself is inherited:
{
"Drivers": [
{
"Driver_Type": "X_SHRED",
"Driving_Allele": "Ad",
"Driving_Allele_Params": {
"Allele_To_Copy": "Ad",
"Allele_To_Replace": "Aw",
"Copy_To_Likelihood": [
{"Copy_To_Allele": "Ad", "Likelihood": 1.0},
{"Copy_To_Allele": "Aw", "Likelihood": 0.0}
]
},
"Shredding_Alleles": {
"Allele_Required": "Yw",
"Allele_To_Shred": "Xw",
"Allele_To_Shred_To": "Xm",
"Allele_Shredding_Fraction": 1.0,
"Allele_To_Shred_To_Surviving_Fraction": 0.0
}
}
]
}
With Allele_Shredding_Fraction = 1.0 and Allele_To_Shred_To_Surviving_Fraction = 0.0,
all X-bearing sperm are destroyed and none survive as the Xm allele.
Y_SHRED example:
A Y-shredding drive that biases offspring toward females. When a male carrying Ad also has
the X-chromosome allele Xw (Allele_Required), his Y-bearing sperm carrying Yw
(Allele_To_Shred) are destroyed:
{
"Drivers": [
{
"Driver_Type": "Y_SHRED",
"Driving_Allele": "Ad",
"Driving_Allele_Params": {
"Allele_To_Copy": "Ad",
"Allele_To_Replace": "Aw",
"Copy_To_Likelihood": [
{"Copy_To_Allele": "Ad", "Likelihood": 1.0},
{"Copy_To_Allele": "Aw", "Likelihood": 0.0}
]
},
"Shredding_Alleles": {
"Allele_Required": "Xw",
"Allele_To_Shred": "Yw",
"Allele_To_Shred_To": "Ym",
"Allele_Shredding_Fraction": 1.0,
"Allele_To_Shred_To_Surviving_Fraction": 0.0
}
}
]
}
Configuration parameters
Gene drivers are defined in the Drivers array within each species entry in
Vector_Species_Params. The following table lists all parameters. Parameters marked as applying
to specific driver types are ignored for other types.
| Parameter | Type | Min | Max | Default | Description |
|---|---|---|---|---|---|
| Drivers | array of json objects | NA | NA | [] | Array of gene driver objects defined per species under Vector_Species_Params. Each entry specifies a drive type and the alleles it acts on. All drivers in the array must be of the same Driver_Type. See example. |
| Driver_Type | string | NA | NA | NA | The type of gene drive mechanism. CLASSIC — the driver copies itself only when one gamete has the driving allele and the other has the target allele to replace. INTEGRAL_AUTONOMOUS — at least one gamete must carry the driver; effectors can be driven even if the driver itself fails to copy. DAISY_CHAIN — a multi-element system where each component drives the next but the first element is not driven, making the system self-limiting. X_SHRED — destroys X-bearing sperm to bias offspring toward males. Y_SHRED — destroys Y-bearing sperm to bias offspring toward females. |
| Driving_Allele | string | NA | NA | NA | The allele that acts as the driver. Must be defined in the species Genes configuration. For X_SHRED and Y_SHRED, this is the allele at a non-gender locus that activates shredding. |
| Alleles_Driven | array of json objects | NA | NA | [] | Defines the alleles to be copied and the probability distribution of outcomes at each driven locus. Used by CLASSIC, INTEGRAL_AUTONOMOUS, and DAISY_CHAIN drive types. Each entry specifies one locus to drive via Allele_To_Copy, Allele_To_Replace, and Copy_To_Likelihood. See example. |
| Allele_To_Copy | string | NA | NA | NA | The allele to be copied into the target gamete. Must be from the same gene/locus as Allele_To_Replace. |
| Allele_To_Replace | string | NA | NA | NA | The allele that must be present in the target gamete and will be replaced. Must be from the same gene/locus as Allele_To_Copy. Must have an entry in Copy_To_Likelihood (representing the failure probability, which may be zero). |
| Copy_To_Likelihood | array of json objects | NA | NA | NA | A list of allele-to-likelihood pairs defining the probability distribution of outcomes when the drive attempts to copy. Entries must include the Allele_To_Replace (probability of copy failure) and sum to 1.0. Additional entries represent successful copy or mutation products. See example. |
| Copy_To_Allele | string | NA | NA | NA | The name of the allele produced at the cut site. Must be from the same gene as Allele_To_Copy and Allele_To_Replace. |
| Likelihood | float | 0 | 1 | 0 | The probability of producing this allele at the cut site. All entries in Copy_To_Likelihood must sum to 1.0. |
| Driving_Allele_Params | json object | NA | NA | NA | For X_SHRED and Y_SHRED only. Defines how the driving allele itself is inherited using the same Allele_To_Copy, Allele_To_Replace, and Copy_To_Likelihood structure as entries in Alleles_Driven. See example. |
| Shredding_Alleles | json object | NA | NA | NA | For X_SHRED and Y_SHRED only. Defines which gender alleles are targeted for destruction during spermatogenesis and how surviving gametes are treated. See example. |
| Allele_Required | string | NA | NA | NA | The gender allele the male must carry for shredding to occur. For X_SHRED, must be a Y-chromosome allele. For Y_SHRED, must be an X-chromosome allele. |
| Allele_To_Shred | string | NA | NA | NA | The gender allele targeted for destruction. For X_SHRED, must be an X-chromosome allele. For Y_SHRED, must be a Y-chromosome allele. |
| Allele_To_Shred_To | string | NA | NA | NA | The allele that shredded gametes are converted to before being eliminated or surviving. Typically a mutant or resistance allele. Must differ from Allele_Required and Allele_To_Shred. |
| Allele_Shredding_Fraction | float | 0 | 1 | 1.0 | The fraction of targeted gametes (Allele_To_Shred) that are converted to Allele_To_Shred_To. A value of 1.0 means all targeted gametes are shredded; less than 1.0 means some survive unchanged. |
| Allele_To_Shred_To_Surviving_Fraction | float | 0 | 1 | 0.0 | The fraction of converted gametes (Allele_To_Shred_To) that survive as viable eggs. 0.0 means perfect shredding — none survive. 1.0 means all converted gametes survive and can contribute to offspring. |