Microsporidia Infection Model
The model extends the standard VECTOR_SIM model to track endosymbiont infections and their effects in mosquito vectors throughout the complete lifecycle — from larval infections through adult transmission events to vertical transmission to offspring.
Although this model was originally created to study Microsporidia MB and its impact on malaria transmission dynamics, it can represent any endosymbiont that spreads between vectors horizontally (during mating) or vertically (from parent to offspring) and reduces the vector's ability to acquire or transmit disease. The framework is not limited to microsporidia — it can model Wolbachia-like symbionts, engineered bacteria, or other intracellular organisms with similar transmission and disease-blocking characteristics.
In the following sections, we will refer to the modeled endosymbionts as "microsporidia" for simplicity, and vector-borne diseases as "malaria", but the framework applies broadly to any vector-endosymbiont-disease system with similar dynamics.
The microsporidia model implements four core transmission processes:
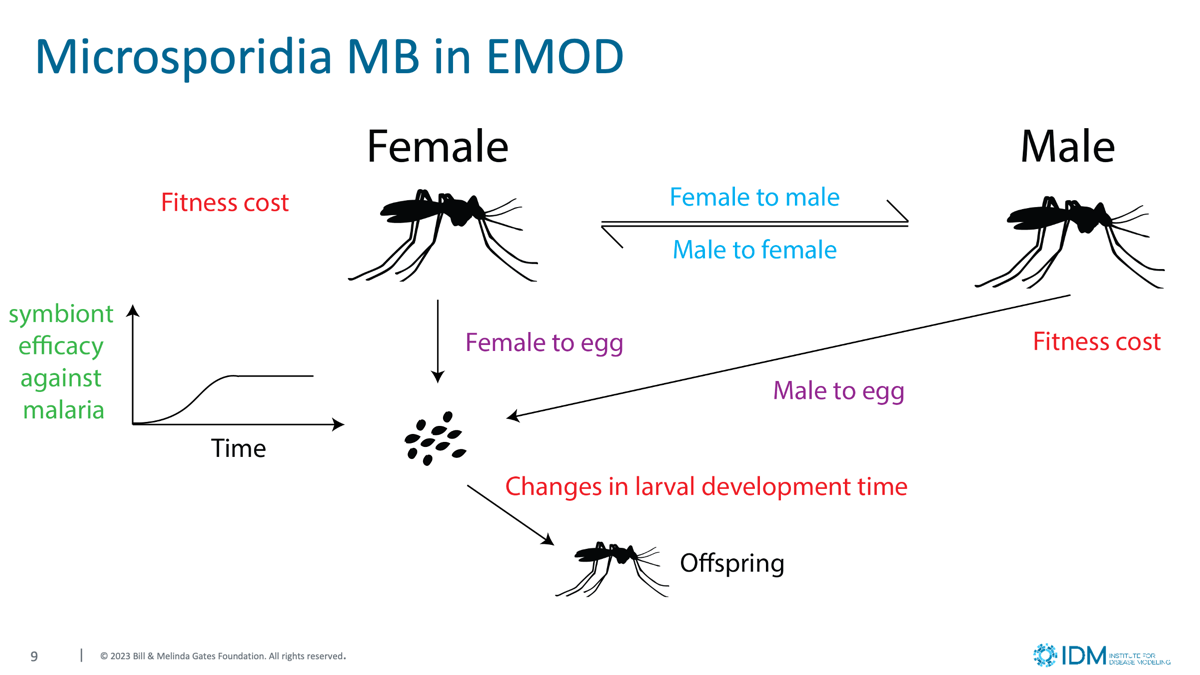
Microsporidia transmission and disease interference in the vector lifecycle
- Vertical transmission: Infected parents transmit microsporidia to their offspring through both maternal (female-to-egg) and paternal (male-to-egg) routes with strain-specific probabilities.
- Horizontal transmission: Adult-to-adult transmission occurs during mating, with infected males and females transmitting their strain to uninfected partners at configurable rates.
- Multi-strain dynamics: The model supports up to 3 distinct microsporidia strains per mosquito species, each with independent transmission parameters and phenotypic effects.
- Malaria interference: Microsporidia infection modifies mosquito susceptibility to malaria acquisition and transmission through time-dependent immunity effects.
Some of the model limitations include:
- Maximum strain count: Limited to 3 microsporidia strains per species.
- Fixed transmission probabilities: Transmission rates are constant and do not vary with environmental conditions or vector age
- No co-infection dynamics: Each vector can carry only one microsporidia strain; competitive exclusion is absolute
- Simplified immunity: Malaria interference effects are time-dependent but do not account for dose-response relationships or microsporidia load
What are microsporidia?
Microsporidia are a diverse group of obligate intracellular eukaryotic parasites that infect a wide range of hosts, including insects. In mosquitoes, certain microsporidia species (such as Microsporidia MB) have been identified as naturally occurring symbionts that can dramatically reduce malaria transmission. These parasites are particularly attractive for biocontrol applications because they:
- Are vertically transmitted from parents to offspring, ensuring persistence in mosquito populations
- Significantly impair Plasmodium development and transmission without severely compromising mosquito fitness
- Can be artificially introduced into wild mosquito populations through targeted release programs
- Provide a transmission-blocking mechanism that complements other malaria control strategies
Field studies have shown that microsporidia-infected Anopheles mosquitoes are significantly less likely to become infected with P. falciparum and, when infected, rarely develop sporozoites capable of transmission. This makes microsporidia an important tool for understanding and potentially controlling malaria transmission dynamics.
Why model microsporidia?
Modeling microsporidia infections is essential for:
Evaluating biocontrol strategies: Microsporidia-based interventions could involve releasing infected mosquitoes to establish symbiont infections in wild populations. Models help predict the conditions under which such releases would be successful and sustainable. Optimizing intervention timing and coverage: The model can evaluate how release timing, coverage levels, and strain characteristics affect the establishment and persistence of transmission-blocking microsporidia. Assessing combined intervention effects: Microsporidia work synergistically with other vector control measures. Models help predict how microsporidia-based interventions interact with insecticides, bed nets, and other malaria control tools. Understanding natural transmission dynamics: Wild mosquito populations often harbor microsporidia at moderate prevalences. Understanding how these infections spread and persist helps explain natural variation in malaria transmission intensity and seasonal patterns. Strain selection and optimization: By comparing different microsporidia strains with varying transmission characteristics and phenotypic effects, models can guide the selection of optimal strains for field deployment.
Transmission mechanisms
Inter-population transmission
When vector migration is used, microsporidia infections migrate with their hosts, enabling spatial spread modeling between different vector populations and metapopulation analysis. The model does not currently include environmental transmission pathways (e.g., spore contamination of larval habitats) beyond the LarvalMicrosporidiaIntervention, see larval-microsporidia-intervention below for details.
Intra-population transmission
The model implements three distinct within population transmission pathways that operate according to empirically derived probabilities:
Adult-to-adult
During mating, microsporidia can transmit between adult mosquitoes through two pathways:
Male-to-female transmission: When an infected male mates with an uninfected female, transmission occurs with probability set by the Male_To_Female_Transmission_Probability parameter.
Female-to-male transmission: When an infected female mates with an uninfected male, transmission occurs with probability set by the Female_To_Male_Transmission_Probability parameter.
If both partners are already infected (regardless of strain), no additional transmission occurs.
Adult-to-egg
Microsporidia transmit from infected parents to their offspring through vertical transmission pathways:
Female-to-egg transmission: Infected females transmit microsporidia to their eggs with probability defined by the Female_To_Egg_Transmission_Probability parameter.
Male-to-egg transmission: Males can also transmit microsporidia to eggs through infected sperm with probability defined by the Male_To_Egg_Transmission_Probability parameter, independent of female infection status.
When both parents are infected, the combined transmission probability is calculated as:
For eggs that become infected through this combined mechanism, the strain is determined by the relative contribution of each parent:
The model uses binomial approximation to determine how many eggs become infected and which strain they carry.
Larval habitat seeding
Microsporidia can also be introduced directly into larval habitats through the LarvalMicrosporidiaIntervention. This intervention mimics environmental seeding of water bodies with microsporidia spores, allowing larvae to become infected through ingestion or contact with contaminated water. See larval-microsporidia-intervention below for details.
Phenotypic effects
Microsporidia infection affects multiple aspects of mosquito biology, with effects varying by strain:
Larval development
Growth rate modification: The Larval_Growth_Modifier parameter adjusts the daily larval growth rate. Values greater than 1.0 accelerate development (as observed with Microsporidia MB), while values less than 1.0 slow development.
Adult mortality
Female mortality effects: The Female_Mortality_Modifier adjusts female adult mortality rates. Values greater than 1.0 increase mortality (shorter lifespan), while values less than 1.0 decrease mortality.
Male mortality effects: The Male_Mortality_Modifier similarly adjusts male adult mortality rates independently from female effects.
Laboratory studies with Microsporidia MB have shown minimal impact on adult survival, but the model allows for strain-specific effects that may differ from field observations.
Malaria transmission interference
Microsporidia's primary value for malaria control lies in their ability to interfere with Plasmodium development and transmission:
Malaria Acquisition interference: The Duration_To_Disease_Acquisition_Modification parameter defines how microsporidia infection affects the probability that a female mosquito becomes infected with malaria when feeding on an infectious human. This is specified as a time-dependent modifier using an dictionary with Times and Values keys, where:
- Times: Array of days since microsporidia infection (ascending order)
- Values: Array of probability multipliers (0-1) for malaria acquisition
Malaria Transmission interference: The Duration_To_Disease_Transmission_Modification parameter similarly affects the probability that an infected mosquito successfully transmits malaria when biting a susceptible human. This captures scenarios where mosquitoes acquire malaria before becoming infected with microsporidia.
- Times: Array of days since microsporidia infection (ascending order)
- Values: Array of probability multipliers (0-1) for malaria transmission
For example, newly infected mosquitoes might show no acquisition or transmission interference (multiplier = 1.0), but interference increases over time as the microsporidia establish infection, reaching maximum effect (multiplier = 0.0) after several days. The model uses linear interpolation to calculate values for times not explicitly defined, with times greater than maximum time defined maintaining the last value of the map.
Configuration parameters
Microsporidia strains are configured within the Vector_Species_Params section as an array called Microsporidia. Each array element defines one strain with the following parameters.
Note
Parameters are case-sensitive. For Boolean parameters, set to 1 for true or 0 for false. Minimum, maximum, or default values of "NA" indicate that those values are not applicable for that parameter.
EMOD does not use true defaults; that is, if the dependency relationships indicate that a parameter is required, you must supply a value for it. However, many of the tools used to work with EMOD will use the default values provided below.
JSON format does not permit comments, but you can add "dummy" parameters to add contextual information to your files. Any keys that are not EMOD parameter names will be ignored by the model.
| Parameter | Type | Min | Max | Default | Description |
|---|---|---|---|---|---|
| Strain_Name | string | NA | NA | "" | A required unique identifier for the microsporidia strain. Must be unique across all species. Cannot be empty or 'NoMicrosporidia'. |
| Female_To_Egg_Transmission_Probability | float | 0 | 1 | 0 | Probability that an infected female transmits microsporidia to her eggs during oviposition. |
| Male_To_Egg_Transmission_Probability | float | 0 | 1 | 0 | Probability that a female mated with an infected male transmits microsporidia to her eggs, independent of her own infection status. |
| Female_To_Male_Transmission_Probability | float | 0 | 1 | 0 | Probability that an infected female transmits microsporidia to an uninfected male during mating. |
| Male_To_Female_Transmission_Probability | float | 0 | 1 | 0 | Probability that an infected male transmits microsporidia to an uninfected female during mating. |
| Larval_Growth_Modifier | float | 0 | 3.40E+38 | 1 | Multiplier for daily larval growth rate for larva infected with this particular strain. Values greater than 1.0 accelerate development; values less than 1.0 slow development. |
| Female_Mortality_Modifier | float | 0 | 3.40E+38 | 1 | Multiplier for female adult mortality rate for vectors infected with this particular strain. Values greater than 1.0 increase mortality (shorter lifespan); values less than 1.0 decrease mortality. |
| Male_Mortality_Modifier | float | 0 | 3.40E+38 | 1 | Multiplier for male adult mortality rate for vectors infected with this particular strain. Values greater than 1.0 increase mortality (shorter lifespan); values less than 1.0 decrease mortality. |
| Duration_To_Disease_Acquisition_Modification | JSON object | NA | NA | NA | Time-dependent modifier for disease acquisition probability for vectors infected with this particular strain. It is a dictionary of arrays where Times represent days since microsporidia infection (ascending order) and Values represent probability multipliers (0-1). The model uses linear interpolation for times not explicitly defined; times beyond the maximum retain the last value. See example. |
| Duration_To_Disease_Transmission_Modification | JSON object | NA | NA | NA | Time-dependent modifier for disease transmission probability for vectors infected with this particular strain. It is a dictionary of arrays where Times represent days since microsporidia infection (ascending order) and Values represent probability multipliers (0-1). The model uses linear interpolation for times not explicitly defined; times beyond the maximum retain the last value. See example. |
The following example shows a two-strain microsporidia configuration for Anopheles gambiae.
{
"Vector_Species_Params": [
{
"Name": "gambiae",
"Microsporidia": [
{
"Strain_Name": "Strain_A",
"Female_To_Egg_Transmission_Probability": 0.7,
"Male_To_Egg_Transmission_Probability": 0.0,
"Female_To_Male_Transmission_Probability": 0.4,
"Male_To_Female_Transmission_Probability": 0.98,
"Larval_Growth_Modifier": 1.08,
"Female_Mortality_Modifier": 1.2,
"Male_Mortality_Modifier": 1.2,
"Duration_To_Disease_Acquisition_Modification": {
"Times": [0, 1, 2, 3],
"Values": [1.0, 0.8, 0.3, 0.0]
},
"Duration_To_Disease_Transmission_Modification": {
"Times": [0, 3, 7],
"Values": [1.0, 0.5, 0.1]
}
},
{
"Strain_Name": "Strain_B",
"Female_To_Egg_Transmission_Probability": 0.5,
"Male_To_Egg_Transmission_Probability": 1.0,
"Female_To_Male_Transmission_Probability": 0.0,
"Male_To_Female_Transmission_Probability": 0.0,
"Larval_Growth_Modifier": 1.0,
"Female_Mortality_Modifier": 1.0,
"Male_Mortality_Modifier": 1.0,
"Duration_To_Disease_Acquisition_Modification": {
"Times": [0, 6],
"Values": [1.0, 0.5]
},
"Duration_To_Disease_Transmission_Modification": {
"Times": [0, 7],
"Values": [1.0, 0.7]
}
}
]
}
]
}
Interventions
LarvalMicrosporidiaIntervention
The LarvalMicrosporidiaIntervention provides a mechanism for introducing microsporidia into larval habitats, simulating environmental release programs or natural habitat contamination. This node-level intervention can target specific habitat types and applies configurable infectivity that may decay over time.
See parameter-campaign-node-larvalmicrosporidiaintervention for details.
MosquitoRelease intervention
Microsporidia can also be introduced through the MosquitoRelease intervention by specifying the Released_Microsporidia_Strain parameter with the desired strain name. This allows for targeted release of infected adult mosquitoes to establish microsporidia in wild populations.
See parameter-campaign-node-mosquitorelease for details.
Output and reporting
The microsporidia model extends several existing reports and adds a dedicated report to track infection dynamics:
-
software-report-vector-stats — When
Include_Microsporidia_Columnsis enabled, adds columns partitioning vector counts by microsporidia status (HasMicrosporidia-STATE_XXX/NoMicrosporidia-STATE_XXX), providing temporal and spatial tracking of how microsporidia infections spread through vector populations. -
software-report-microsporidia — A specialized report focusing exclusively on microsporidia dynamics, tracking infection prevalence by strain across vector life stages and monitoring strain competition dynamics.
When vector migration is enabled, microsporidia infections migrate with their hosts, enabling spatial spread modeling and metapopulation analysis.
Implementation notes
Performance considerations
The microsporidia model adds computational overhead through:
- Adult mating algorithms: Pairing males and females for transmission calculations requires more complex mating logic than the base model
- Multi-strain tracking: Maintaining separate cohorts for different microsporidia strains increases memory usage and computational complexity
- Time-dependent effects: Evaluating time-since-infection modifiers for disease interference requires additional calculations per vector per timestep