Vector genetics
Vector control interventions such as insecticide-treated nets (ITNs) and indoor residual spraying (IRS) are cornerstones of malaria control and elimination efforts. However, the growing spread of insecticide resistance in mosquito populations threatens to undermine these tools. In parallel, new strategies such as gene drives — genetic elements that bias their own inheritance to spread through a population within a few generations — have been proposed as a way to either suppress vector populations or replace them with vectors that are unable to transmit malaria.
EMOD's vector genetics system provides a stochastic, agent-based framework for simulating the inheritance and phenotypic effects of genetic loci in mosquito populations. Each mosquito carries a diploid genome composed of up to 9 genetic loci with up to 8 alleles per locus. The system supports Mendelian inheritance, germline mutations, insecticide resistance, gene drives, sex-ratio distortion, and releases of genetically modified mosquitoes — all operating within the standard vector lifecycle described in vector-model-transmission. This allows researchers to investigate the spread of insecticide resistance, evaluate gene drive deployment strategies, and explore the interaction between vector genetics and malaria transmission in a spatially and temporally explicit setting.
For more details on the modeling approach and example applications, see Selvaraj et al. (2020), Vector genetics, insecticide resistance and gene drives: An agent-based modeling approach to evaluate malaria transmission and elimination.
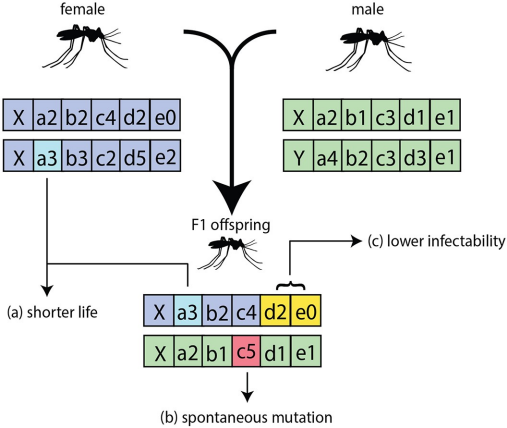
Overview of the vector genetics system in EMOD: a female and male each carry a diploid genome; the F1 offspring inherits one gamete from each parent. The labels illustrate three consequences modeled by the genetics system: (a) allele combinations can modify phenotypic traits such as lifespan (see trait-modifiers), (b) spontaneous germline mutations can introduce new allele variants (see germline-mutations), and (c) allele combinations can alter vector competence such as infectability (see trait-modifiers). From Selvaraj et al. (2020), doi:10.1371/journal.pcbi.1008121.
Vector genetics is configured per species under Vector_Species_Params in config.json. Each
species defines its own set of genes, alleles, trait modifiers, and (optionally) gene drives.
The system does not require a separate simulation type — it is available whenever the simulation
includes vector population dynamics (Simulation_Type set to VECTOR_SIM or MALARIA_SIM).
Genome representation
In EMOD, each mosquito carries a simplified diploid genome composed of two gametes — one inherited from each parent. The genome supports up to 8 user-defined genetic loci, each with up to 8 named alleles. While real mosquito genomes contain thousands of genes, this abstraction captures the loci most relevant to the dynamics being studied. A mosquito's Wolbachia status and microsporidia strain are also tracked per individual alongside the genetic information. Each female also stores her mate's genome for the duration of her life — represented in the VectorCohort as the "Mate" genome alongside her own ("Self") — and uses it when she lays eggs after each feeding cycle.
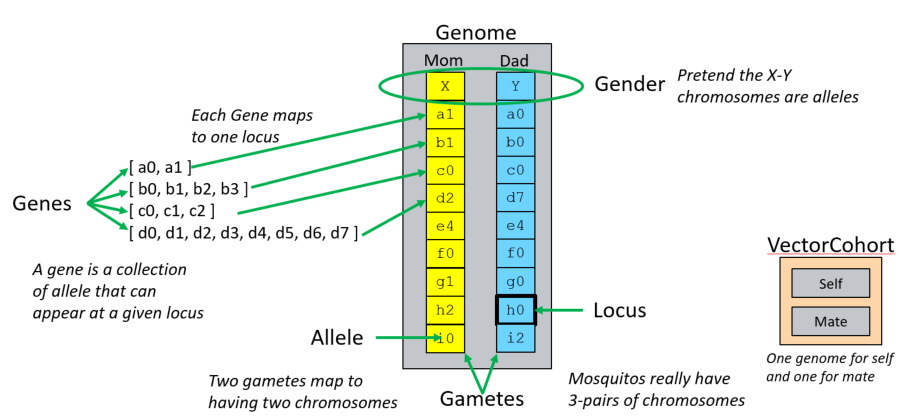
EMOD genome representation: the genome consists of two gametes (Mom and Dad), each carrying one allele per locus. The top row is the gender locus, where X and Y are modeled as alleles. The remaining rows are user-defined loci; the possible alleles for each gene are listed on the left. At right, the VectorCohort structure shows that each female stores two genomes — one for herself (Self) and one for her mate (Mate) — used during fertilization.
One locus is always reserved as the gender gene. A mosquito's sex is always determined by the gender gene — this is true even in simulations that do not use the genetics system. In a standard simulation the gender gene has just two alleles (X and Y), but it can be configured with additional alleles to support gene drive scenarios such as sex-ratio distortion, where a drive element on the X chromosome suppresses Y-bearing sperm.
Genes and alleles
Genes are defined in the Genes array within each species' configuration. Each gene declares a
set of named alleles with initial population frequencies to be used at simulation initialization.
{
"Vector_Species_Params": [
{
"Name": "gambiae",
"Genes": [
{
"Is_Gender_Gene": 1,
"Alleles": [
{"Name": "X", "Initial_Allele_Frequency": 0.75, "Is_Y_Chromosome": 0},
{"Name": "Y", "Initial_Allele_Frequency": 0.25, "Is_Y_Chromosome": 1}
]
},
{
"Alleles": [
{"Name": "a0", "Initial_Allele_Frequency": 0.9, "Is_Y_Chromosome": 0},
{"Name": "a1", "Initial_Allele_Frequency": 0.1, "Is_Y_Chromosome": 0}
]
}
]
}
]
}
Gender gene: If defined, the gender gene must be listed first. It partitions alleles into
X-chromosome alleles (Is_Y_Chromosome = 0 (false)) and Y-chromosome alleles
(Is_Y_Chromosome = 1 (true)). If no gender gene is explicitly defined, EMOD creates a default
one with a single X allele and a single Y allele, with XX and XY combinations being generated at
about 50/50 frequencies.
Constraints: Allele names must be unique across all genes within a species. Frequencies within a gene must sum to 1.0. A species may define at most 9 genes (including the gender gene) and at most 8 alleles per gene.
When the simulation initializes, the vector population is seeded with genomes drawn from the configured allele frequencies. Each gamete's alleles are sampled independently at each locus, then maternal and paternal gametes are combined to produce diploid genomes. The resulting genotype frequencies follow Hardy-Weinberg expectations.
Genetic diversity
Keep the following in mind when deciding how much genetic diversity to define for your vector population:
-
Initial population size: If you want high genetic diversity in the initial population, the initial adult population must be large enough to represent that diversity. For example, if you define enough genes and alleles to produce 100,000 possible genome combinations but only initialize 10,000 mosquitoes, many combinations will simply not be present. Population size also needs to be sufficient that stochastic effects such as daily mortality do not eliminate rare alleles before the population becomes established.
-
Runtime cost: Greater genetic diversity — even if introduced later via a release — increases simulation run time. More distinct genomes mean more objects to track and process at each time step.
Mating
In EMOD, alleles are spread through the population via mating. The model simplifies real mosquito mating behavior: females mate exactly once in their lifetime, when they transition from immature to adult, and males are available for mating every day. In reality, mating dynamics are more complex, but this approximation captures the key genetic consequences — each female's offspring share a single father. During the immature-to-adult transition, immature males are updated first to determine how many of each genome are available in the male queue. Female cohorts then select mates randomly from the available males, weighted by population count.
Each female stores her mate's genome for the duration of her life. When she completes a feeding cycle and is ready to oviposit, the stored mate genome is used to determine offspring genotypes through the fertilization process described below.
Mendelian inheritance
In EMOD's implementation of fertilization, each parent contributes one gamete to the offspring
through standard Mendelian segregation: at each locus, one of the two parental alleles is
selected with equal (50/50) probability, and loci segregate independently. The model assumes
all loci are on separate chromosomes (no linkage), which is a simplification of real genetics
where nearby loci on the same chromosome tend to be inherited together. Females always contribute an X-bearing
gamete; males contribute either X or Y, with the ratio controlled by the FEMALE_EGG_RATIO
trait modifier. All possible gamete combinations are enumerated, each assigned a probability
equal to the product of the two gamete probabilities, and eggs are distributed across these
combinations stochastically. Offspring sharing the same genome are grouped into cohorts and
enter the standard egg-to-adult development pipeline.
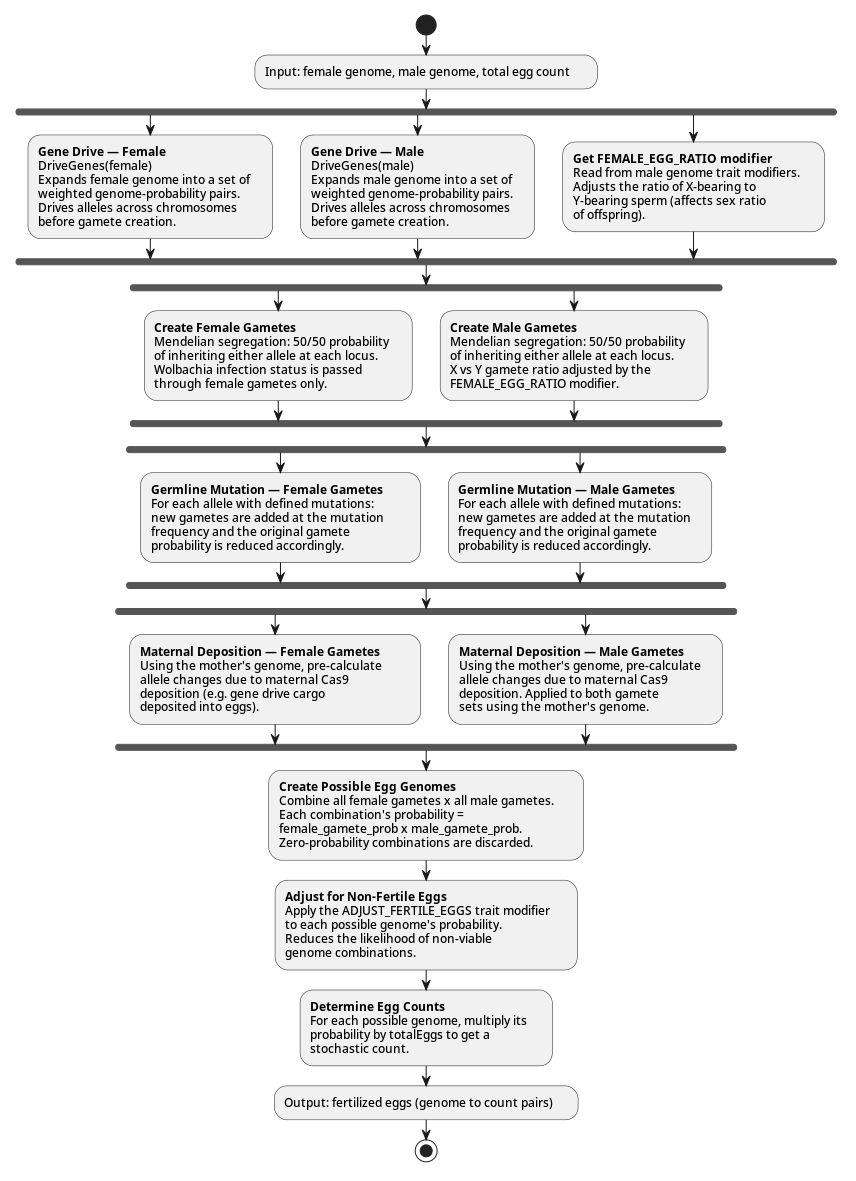
The following diagram shows the full sequence of events during fertilization, including when gene drive, germline mutation, and maternal deposition are applied relative to gamete creation:
Germline mutations
In EMOD, alleles can mutate during gametogenesis, producing new allele variants in offspring at a configured per-generation rate. This is a simplified model of mutation — real mutation processes involve a variety of mechanisms (point mutations, insertions, deletions, etc.) that are abstracted here into a single probability of one named allele converting to another. Mutations are defined per gene:
{
"Genes": [
{
"Alleles": [
{"Name": "a0", "Initial_Allele_Frequency": 0.95},
{"Name": "a1", "Initial_Allele_Frequency": 0.05}
],
"Mutations": [
{
"Mutate_From": "a0",
"Mutate_To": "a1",
"Probability_Of_Mutation": 0.005
}
]
}
]
}
When gametes are created, each allele has the configured probability of mutating to the specified target. The mutation is applied after standard Mendelian segregation but before fertilization. Multiple mutations can be defined for a single gene, including bidirectional mutations.
Note that germline mutation acts on gametes, not the parent genome. New mutant gametes are added at the defined frequency, and the probability of the original (unmutated) gamete is reduced by the same amount, so total probability is conserved.
Trait modifiers
In EMOD, trait modifiers are the mechanism by which genotype influences mosquito biology. They map allele combinations to phenotypic effects — controlling traits such as mortality, fecundity, insecticide susceptibility, and parasite transmission. Real genotype-phenotype relationships are far more complex, involving epistasis, environmental interactions, and polygenic effects; EMOD simplifies this to direct multipliers on specific traits for specified allele combinations.
Each modifier specifies one or more Allele_Combinations (the genotypes it applies to) and one
or more Trait_Modifiers (the traits it affects and by how much):
{
"Gene_To_Trait_Modifiers": [
{
"Allele_Combinations": [["a1", "a1"]],
"Trait_Modifiers": [
{"Trait": "MORTALITY", "Modifier": 1.5}
]
}
]
}
In this example, mosquitoes homozygous for a1 at a given locus have 1.5x the baseline
mortality rate.
Allele combination syntax: Each combination is a list of allele names — two per locus (one for
each gamete). "*" acts as a wildcard matching any allele. For a gene with alleles a0 and
a1:
["a1", "a1"]— matches onlya1/a1homozygotes["a1", "*"]— matches any mosquito carrying at least onea1allele["*", "*"]— matches all genotypes (wildcard at both positions)
Multi-locus combinations list allele pairs for each relevant locus. Loci not mentioned are treated as wildcards.
When multiple modifiers apply to the same trait for a given genome, their values are multiplied together.
Available traits
The following traits can be modified by genotype:
INFECTED_BY_HUMAN, 1.0, "Multiplier on the probability that a mosquito becomes infected when feeding on an infectious human. Applied to the species-level ``Acquire_Modifier`` parameter."
FECUNDITY, 1.0, "Multiplier on the number of eggs laid per oviposition. Applied to the ``Egg_Batch_Size`` parameter. This impacts egg count before egg crowding takes effect."
FEMALE_EGG_RATIO, 1.0, "Controls the sex ratio of offspring. A value of 1.0 produces 50/50 male/female. Values above 1.0 bias toward female; values below 1.0 bias toward male. At 2.0 all offspring are female; at 0.0 all are male. Applied during fertilization after egg crowding."
STERILITY, 1.0, "Determines if eggs are viable based on the parents' genomes. A value of 0.0 means the vector is sterile — if either parent is sterile, the eggs are not viable and are not added to the egg queue. Any nonzero value means fertile. Applied after egg crowding. Sterility does not impact mating. If the female mates with a sterile male, then she will feed as normal but produce no eggs."
TRANSMISSION_TO_HUMAN, 1.0, "Multiplier on the probability that sporozoites in the salivary gland successfully infect a human during a bite. Applied to the species-level ``Transmission_Rate`` parameter."
ADJUST_FERTILE_EGGS, 1.0, "Multiplier on the probability of each genome's eggs being fertile. This is the last step in the fertilization process before actual numbers of eggs are assigned. A value of 0.0 means no eggs are produced; 1.0 means no change; values above 1.0 increase egg production for that genome."
MORTALITY, 1.0, "Multiplier on the daily mortality rate, where the base rate is ``1/Adult_Life_Expectancy`` (or ``1/Male_Life_Expectancy`` for males). Values above 1.0 increase mortality (shorter lifespan); values below 1.0 decrease it."
INFECTED_PROGRESS, 1.0, "Multiplier on the daily progression from infected to infectious (oocyst-to-sporozoite conversion rate)."
OOCYST_PROGRESSION, 1.0, "Additional multiplier on temperature-dependent oocyst maturation. Only applies when the vector carries a parasite matching a specified barcode (requires Full Parasite Genetics). This is an additional multiplier on top of INFECTED_PROGRESS."
SPOROZOITE_MORTALITY, 1.0, "Multiplier on sporozoite death rate within the vector. Only applies when the vector carries a parasite matching a specified barcode (requires Full Parasite Genetics)."
Insecticide resistance
Insecticide resistance is modeled through the interaction of vector genotype with insecticide properties. Each insecticide in the simulation defines resistance profiles that specify which allele combinations confer resistance and how much protection they provide.
Please see vector-model-insecticide-resistance for more information on configuring insecticide resistance.
Gene drives
Gene drives are genetic elements that bias their own inheritance, spreading through a population at rates exceeding standard Mendelian expectations. EMOD supports five gene drive types, each modeling a different mechanism.
Note that gene drive is applied before gamete creation. A single parent genome is first expanded into a set of weighted genome-probability pairs by the drive, and Mendelian segregation then operates on those possibilities.
Please see vector-model-gene-drives for more information on configuring gene drives.
Maternal deposition
Maternal deposition models the transfer of Cas9 protein (not DNA) from mother to offspring,
generating resistance alleles in the embryo before the drive's homology-directed repair is
active. It extends gene drive behavior by adding a pre-embryonic cutting step configured in the
Maternal_Deposition array within Vector_Species_Params.
Note that maternal deposition uses the mother's genome to modify both the female and male gamete sets. The Cas9-driven allele changes originate from the mother regardless of which set of gametes is being modified.
Please see vector-model-maternal-deposition for full details on how maternal deposition works, configuration parameters, and validation rules.
Wolbachia
Wolbachia is an intracellular bacterium found naturally in many insect species and can be introduced into mosquito populations as a disease-control strategy. In Anopheles mosquitoes, Wolbachia infection can shorten adult lifespan — important because malaria parasites (Plasmodium) require 10–14 days to develop inside the mosquito before it can transmit disease, so a shorter lifespan means fewer mosquitoes survive long enough to become infectious. Wolbachia can also inhibit Plasmodium development directly by activating the mosquito immune system and competing with the parasite for resources. A key feature of Wolbachia for population-level strategies is cytoplasmic incompatibility: infected males cannot successfully reproduce with uninfected females, which causes Wolbachia to spread through a population over time once introduced. Together, these effects — reduced lifespan, parasite inhibition, and self-sustaining spread — make Wolbachia a candidate tool for reducing malaria transmission without eliminating mosquitoes.
EMOD models Wolbachia as a discrete state per vector with four possible values: none, strain A, strain B, or both strains A and B. This is a simplification — in reality, Wolbachia dynamics involve variable bacterial loads, incomplete maternal transmission, and strain-specific effects that are more nuanced than the model captures.
In EMOD, Wolbachia is inherited maternally with 100% fidelity — infected females pass their Wolbachia status to all offspring through the egg cytoplasm. Males do not transmit Wolbachia.
Wolbachia-infected males are incompatible with uninfected females: matings between Wolbachia- carrying males and uninfected females produce inviable eggs (cytoplasmic incompatibility). This is checked during egg laying, and incompatible crosses produce no eggs.
Wolbachia modifies vector biology through two parameters: Wolbachia_Mortality_Modification
is a multiplier on the mortality rate of infected vectors, and Wolbachia_Infection_Modification
is a multiplier on the probability that a Wolbachia-infected vector acquires a malaria infection
when biting an infectious human. A value below 1.0 reduces susceptibility, modeling the parasite-
blocking effect of Wolbachia; the default of 1.0 means no effect.
Vectors with specific Wolbachia status can be introduced into the population using the
MosquitoRelease intervention with the Released_Wolbachia parameter.
Releasing vectors with specific genomes
The MosquitoRelease intervention introduces vectors with user-specified genomes into the
simulation. This is the primary mechanism for modeling releases of genetically modified
mosquitoes, including gene drive carriers and sterile males. Released vectors can also be given
a specific Wolbachia status (via Released_Wolbachia) or a microsporidia strain (via
Released_Microsporidia_Strain). The user specifies the complete genome of the released
vectors — all genes and loci must be specified, including sex (via the appropriate X/Y
chromosome pair).
When gene drive alleles are released, the drive mechanics take effect during subsequent mating and fertilization events, propagating the driven alleles through the population.
See parameter-campaign-node-mosquitorelease for more information on configuring mosquito releases.
Output
software-report-vector-genetics (ReportVectorGenetics) is the primary report for
tracking vector genetics. It produces a CSV file with vector counts stratified by genome, allele,
or allele frequency at each time step, node, and vector state (eggs, larvae, immature, adult,
infected, infectious, male).